Homework 11: Simple linear regression
Objectives:
- Explore null and alternate hypothesis concepts for coefficient estimates and linear models.
- Evaluate error rates (Type I, Type II) and critical value, p-value.
- To compute how to obtain and interpret linear models in R Commander.
- To compute how to obtain and interpret regression statistics and diagnostic plots in R Commander.
- To apply General Linear Model approach to regression models.
Homework 11 expectations
Read through the entire homework before starting to answer a question — all questions are intended to help you achieve the learning outcomes for the chapter. You are expected to have read the chapter and to have completed preceding homework. You are expected to have read the chapter and to have completed preceding homework. Answers are provided to odd numbered problems — turn in your work for even numbered problems. A BONUS opportunity is also provided.
How to work this homework
This homework is in two parts and involves two separate data sets. First, body mass and brain mass of several mammal species. Second, work with the data set cars, stopping distance by speed of the car, develop simple linear regression (SLR) models with diagnostic graphs.
Your report will consist of your answers to the bold, numbered questions (the even ones!) and supporting statistics from R. Suggested steps in your analysis are provided as numbered items (regular, not bold format). You may work together or individually, but each of your must turn in your own report. Don’t “plagiarize” from each other. Do include in your report who you worked with.
What to turn in: Turn in two properly named pdf files:
- Model 1: body mass predicts brain mass
- SLR
The pdfs file contain relevant R code, statistical results — edited to support your answers to the questions, and your answer to the questions (even numbered only). Use of RMarkdown recommended — because it is a simple way to include graphs generated; however copy/paste into a word document is also acceptable.
Notes. By relevant we mean provide just the R code and results from R functions necessary to support your answers to the questions. For example, do not include
- the entire data set when head(dataset) will do
- screenshots of R output!! R output is text — copy/paste
- all statistical output from an R function.
See Part09: Making a report for an example homework file.
Submit your work to CANVAS. Obey proper file naming formats.
Resources for this homework
Chapter 17. Mike’s Biostatistics Book
Mike’s Workbook for Biostatistics: A quick look at R and R Commander, Part01 – Part10 and previous homework pages presented in this workbook.
Additional R commands and or code provided below.
Work on model 1
Objective 1: To learn how to obtain and interpret linear model in R and R Commande.
Dataset
For correlation, use this dataset = Animals. This data set consists of brain weight and body mass of 28 species, including mammals (24 species plus humans) and three dinosaurs (Brachiosaurus, Dipliodocus, Triceratops). The ratio of brain weight to body mass (brain-body mass ratio) is related to the encephalization quotient (EQ); humans and other primates have a high ratio, so this measure is sometimes taken as a crude estimate of “intelligence.”
Note. My description isn’t technically the definition of EQ: see Jerison 1985 and van Schaik et al 2021.
The data set is one of the built-in datasets with R, go to Rcmdr: Data in packages → Read data set from attached package… Select MASS package, then find Animals). Or, just load the data by submitting the code
data(Animals, package="MASS")
Alternatively, for your convenience, I’ve included the dataset at the end of this page (scroll down or click here).
Note. Since the 1990s, comparative biologists have recognized that such species comparisons without accounting for phylogenetic structure of the data likely violates the assumption of independence among the data points. Many approaches to account for the nonindependence have been published: see Chapter 20.12 in Mike’s Biostatistics Book for an introduction to one method now called phylogenetically independent contrasts (PIC). For the purposes of this homework we ignore this issue.
Questions: The work to do for correlation analysis
1. Make a scatterplot of brain weight on body weight.
a) It would a good idea to highlight specific points in the graph.
Note. “… highlight specific points…” This can be done, but it is an advanced R trick (see ggplot2 and gghighlight package). Here’s one method, a bit crude, but it works. Copy and paste the code into the script window of Rcmdr. Assuming you’ve already loaded the dataset Animals, then submit the following code one line a time
classType <- c("M","M","M","M","M","D","M","M","M","M","M","M","M","H","M","D","M","M","M","M","M","M","M","M","M","M","D","M")
dAnimals <-data.frame(classType,Animals)
scatterplot(brain~body | classType, regLine=FALSE, smooth=FALSE, by.groups=TRUE, pch=c(19,19,19), cex=c(2,2,2), col=c("black", "blue", "red"), grid=FALSE, data=dAnimals)
2. Make histograms of body weight and brain weight
3. Conduct a test of normality of brain weight, and another test for body weight
Rcmdr: Statistics → Summaries → Test of normality (Compare Shapiro Wilks against Anderson-Darling)
R code
RcmdrMisc::normalityTest(~body, test="shapiro.test", data=Animals)
Work on Ordinary Linear (Simple) Regression
Objective 2: To learn how to obtain and interpret regression statistics and diagnostic plots in R Commander
Objective 3: To compare models and evaluate “best fit,” with use of R2, the coefficient of determination.
Objective 4: Introduce the General Linear Model approach
For simple linear regression use this dataset = cars, Speed and Stopping Distances of Cars (one of the built-in datasets with R, go to Rcmdr: Data in packages → Read data set from attached package… Select datasets package, then find cars). More simply, at the R prompt write and submit
data(cars, package="datasets")
Alternatively, and for your convenience, I’ve included the dataset at the end of this page (scroll down or click here). The data set is a classic from the 1920s; it’s about distance (feet) to stop a car by speed (mph) of a car (see Wikipedia). Thus, the cars in 1920s looked more like the one on left than the one at right (Fig 2).

|
|
Figure 2. The stopping distance data set is an old one — data from a car similar to the Ford Model A at left, not the Ford Mustang Mach E GT at right.
The work to do on simple linear regression
1. Identify the dependent and independent variables. Justify your selection
2. Make a scatterplot of Distance on Speed
3. Make a histogram of Distance
4. Conduct a test of normality on Distance
Rcmdr: Statistics → Summaries → Test of normality (Compare Shapiro Wilks against Anderson-Darling)
Question 1. Make a preliminary conclusion about whether or not the data conforms to assumptions of linear regression based on your results from items 1 – 4. Include your justification for choice of independent and dependent variables. If you find the data do not conform, create new log-transformed, log10(), variables and redo your assumption tests.
5. Based on your answer to Question 1, write out the model, e.g., in R, the model is specified as Y ~ X, then go ahead and conduct the linear regression of Distance and Speed.
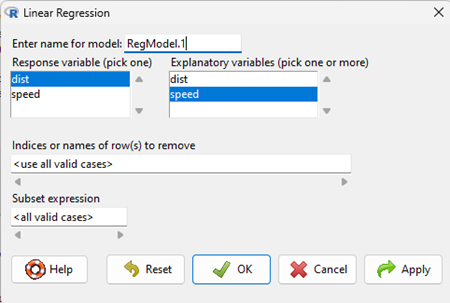
Rcmdr: Statistics → Fit models → Linear model
- an example screen is shown (yours will look different from the example). Note also that you can let Rcmdr assign a name for the model object or you can specify one yourself (⮜ DrD recommended!)
Figure 3. Screenshot linear regression menu R Commander.
R code
RegModel.1 <- lm(dist~speed, data=cars) summary(RegModel.1)
6. Obtain the following from the R output ⮜ recommend you organize these into a table
a) value of the Y-intercept
b) Report on whether the Y-intercept is statistically significant (what is the null hypothesis?)
c) value of the slope
d) Report on whether the slope is statistically significant (what is the null hypothesis?)
e) Write out the statistical model
f) Find the fit statistic for this model.
Question 2. Make a preliminary conclusion about the predictive value of your X (independent) on your Y (dependent) variable
7. Repeat steps 5 and 6, but for the untransformed response variable (– or, if you used the raw, untransformed response variable previously, now run the regression with the transformed variable).
a) record the linear model
b) record the fit statistics (R2 and RSE).
c) compare fit of regressions of the transformed vs untransformed variables. Which is best?
8. Obtain appropriate diagnostic plots and diagnostic statistics for your regression and evaluate your model(s) against the assumptions
Rcmdr: Models → Graphs → Basic diagnostic plots
R code
plot(RegModel.1)
Rcmdr: Models → Numerical diagnostics → “several to choose from” ⮜ part of the evaluation is whether you select the correct diagnostic tests; hint: most of these are listed in your Chapter 18)
R code example, RESET test for nonlinearity
resettest(dist ~ speed, power=2:3, type="regressor", data=cars)
Question 3. Make a conclusion about the regression model based on your results from Question 1 and Question 2.
Question 4. Use your regression model to predict stopping distance at 60 mph. How does your prediction compare to stopping distance of 108 feet for a Ford Mustang at the same speed?
Hint: you can simply calculate this using the appropriate numbers from your regression model. Instead of “hand” calculations, alternatively, you could use the following command
predict(modelName, data.frame(nameOfdependent=c(60)))
where modelName is replaced with the object name for your regression model, and nameOfdependent is replaced with the name of your independent variable.
Regardless of method you choose, if you used log-transform, then don’t forget to report your predicted value in original raw form. For example, if log(x, 10), then the antilog 10^x, with x equal to your predicted value. This is a standard recommendation for data analysis — do your statistics on the transformed data, but for graphics or other reports, always back-transform the data.
Save your Markdown file and include SLR as part of your properly named pdf file name. Submit your file to this page.
This concludes expected work for Homework 11.
Bonus
Remember our M&M counts? Consider a new hypothesis — instead of equal color probabilities, perhaps the counts reflect purpose, to minimize costs. Food color dyes differ, and given the tens of billions of M&Ms produced, even small differences for dye costs add up to real money.
Table 1. Summary table M&Ms.
| Color | Cost | Count |
| Blue | 143 | |
| Yellow | 143 | |
| Red | 76 |
Bonus question 1: Develop a simple linear model of dye cost versus counts of the different M&M beans sampled from our collection of Mini M&M bags. Of course, part of your work should include a relevant plot and you should include both a “science hypothesis” along with an appropriate statistical hypothesis for the regression model.
R or Rcmdr commands
myData <- read.table(header=TRUE, sep="t", text = " insert your data table here ") head(myData)
Test normality.
Rcmdr → Statistics → Summaries → Test for normality
Other R/Rcmdr commands provided in text
Data
Dataset = Animals from R package MASS
| Animal | body | brain |
| Mountain beaver | 1.35 | 8.1 |
| Cow | 465 | 423 |
| Grey wolf | 36.33 | 119.5 |
| Goat | 27.66 | 115 |
| Guinea pig | 1.04 | 5.5 |
| Dipliodocus | 11700 | 50 |
| Asian elephant | 2547 | 4603 |
| Donkey | 187.1 | 419 |
| Horse | 521 | 655 |
| Potar monkey | 10 | 115 |
| Cat | 3.3 | 25.6 |
| Giraffe | 529 | 680 |
| Gorilla | 207 | 406 |
| Human | 62 | 1320 |
| African elephant | 6654 | 5712 |
| Triceratops | 9400 | 70 |
| Rhesus monkey | 6.8 | 179 |
| Kangaroo | 35 | 56 |
| Golden hamster | 0.12 | 1 |
| Mouse | 0.023 | 0.4 |
| Rabbit | 2.5 | 12.1 |
| Sheep | 55.5 | 175 |
| Jaguar | 100 | 157 |
| Chimpanzee | 52.16 | 440 |
| Rat | 0.28 | 1.9 |
| Brachiosaurus | 87000 | 154.5 |
| Mole | 0.122 | 3 |
| Pig | 192 | 180 |
Dataset = cars
| speed | dist |
| 4 | 2 |
| 4 | 10 |
| 7 | 4 |
| 7 | 22 |
| 8 | 16 |
| 9 | 10 |
| 10 | 18 |
| 10 | 26 |
| 10 | 34 |
| 11 | 17 |
| 11 | 28 |
| 12 | 14 |
| 12 | 20 |
| 12 | 24 |
| 12 | 28 |
| 13 | 26 |
| 13 | 34 |
| 13 | 34 |
| 13 | 46 |
| 14 | 26 |
| 14 | 36 |
| 14 | 60 |
| 14 | 80 |
| 15 | 20 |
| 15 | 26 |
| 15 | 54 |
| 16 | 32 |
| 16 | 40 |
| 17 | 32 |
| 17 | 40 |
| 17 | 50 |
| 18 | 42 |
| 18 | 56 |
| 18 | 76 |
| 18 | 84 |
| 19 | 36 |
| 19 | 46 |
| 19 | 68 |
| 20 | 32 |
| 20 | 48 |
| 20 | 52 |
| 20 | 56 |
| 20 | 64 |
| 22 | 66 |
| 23 | 54 |
| 24 | 70 |
| 24 | 92 |
| 24 | 93 |
| 24 | 120 |
| 25 | 85 |
/MD